NEBNext® Enzymatic Methyl-seq Kit
- 价格: ¥10749/盒
- 发布日期: 2023-07-05
- 更新日期: 2026-03-17
产品详请
| 产地 |
|
| 品牌 |
New England Biolabs
|
| 货号 |
E7120S
|
| 用途 |
|
| 包装规格 |
24T
|
| 纯度 |
%
|
| CAS编号 |
|
| 别名 |
|
| 是否进口 |
是
|
View or download extensive performance data in our Technical Note .
The NEBNext Enzymatic Methyl-seq Kit provides a high-performance enzyme-based alternative to bisulfite conversion for methylome analysis using Illumina® sequencing.
Libraries are prepared using as little as 10 ng input DNA and the supplied NEBNext Ultra II reagents and the optimized EM-seq Adaptor. TET2 then oxidizes 5-mC and 5-hmC, providing protection from deamination by APOBEC in the next step. In contrast, unmodified cytosines are deaminated to uracils. Libraries are then amplified using a NEBNext master mix formulation of Q5U® (a modified version of Q5® High-Fidelity DNA Polymerase), and sequenced using Illumina instrumentation.
The consistently high conversion performance and minimized DNA damage with the EM-seq protocol, in combination with highly efficient Ultra II library prep, result in superior detection of CpGs with fewer sequencing reads.
Features:
-
Superior sensitivity of detection of 5-mC and 5-hmC
-
Greater mapping efficiency
-
More uniform GC coverage
-
Detection of more CpGs with fewer sequence reads
-
Uniform dinucleotide distribution
-
High-efficiency library preparation, with larger library insert sizes
-
Conversion module also available separately
Figure 1: NEBNext Enzymatic Methyl-seq and Sodium Bisulfite Conversion Methods
 Figure 2: NEBNext Enzymatic Methyl-seq identifies more CpGs than WGBS, at lower sequencing coverage depth
Figure 2: NEBNext Enzymatic Methyl-seq identifies more CpGs than WGBS, at lower sequencing coverage depth
A.

B.
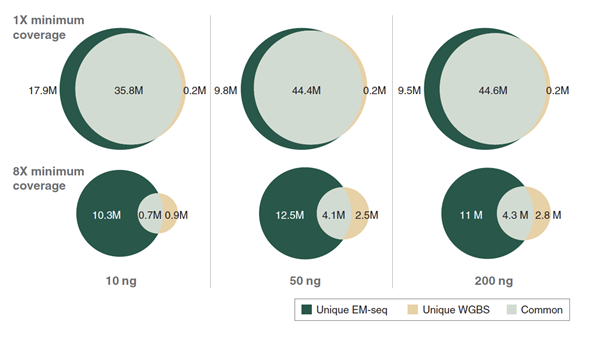 10, 50 and 200 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris® S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold™ Kit for bisulfite conversion. Libraries were sequenced on an Illumina NovaSeq® 6000 (2 x 100 bases). Reads were aligned to hg38 using bwa-meth 0.2.2. Coverage of CpGs with EM-seq and WGBS libraries was analyzed using 324 million paired end reads.
A: Each top and bottom strand CpGs were counted independently, yielding a maximum of 56 million possible CpG sites. EM-seq identifies more CpGs at lower depth of sequencing.
10, 50 and 200 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris® S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold™ Kit for bisulfite conversion. Libraries were sequenced on an Illumina NovaSeq® 6000 (2 x 100 bases). Reads were aligned to hg38 using bwa-meth 0.2.2. Coverage of CpGs with EM-seq and WGBS libraries was analyzed using 324 million paired end reads.
A: Each top and bottom strand CpGs were counted independently, yielding a maximum of 56 million possible CpG sites. EM-seq identifies more CpGs at lower depth of sequencing.
B:The number of unique and common CpGs identified by EM-seq and WGBS at 1x and 8x minimum coverage for each input amount are shown. EM-seq covers at least 20% more CpGs than WGBS at 1x minimum coverage threshold. The difference in CpG coverage increases to two-fold at 8x minimum coverage threshold.
View additional performance data in our
technical note .
Figure 3: NEBNext Enzymatic Methyl-seq has superior uniformity of GC coverage
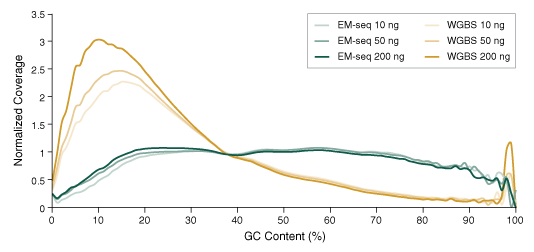 10, 50 and 200 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold Kit for bisulfite conversion. Libraries were sequenced on an Illumina NovaSeq 6000 (2 x 100 bases). Reads were aligned to hg38 using bwa-meth 0.2.2. GC coverage was analyzed using Picard 2.17.2 and the distribution of normalized coverage across different GC contents of the genome (0-100%) was plotted. EM-seq libraries have significantly more uniform GC coverage, and lack the AT over-representation and GC under-representation typical of WGBS libraries.
10, 50 and 200 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold Kit for bisulfite conversion. Libraries were sequenced on an Illumina NovaSeq 6000 (2 x 100 bases). Reads were aligned to hg38 using bwa-meth 0.2.2. GC coverage was analyzed using Picard 2.17.2 and the distribution of normalized coverage across different GC contents of the genome (0-100%) was plotted. EM-seq libraries have significantly more uniform GC coverage, and lack the AT over-representation and GC under-representation typical of WGBS libraries.
View additional performance data in our
technical note .
Figure 4: NEBNext Enzymatic Methyl-seq Libraries have Larger Insert Sizes
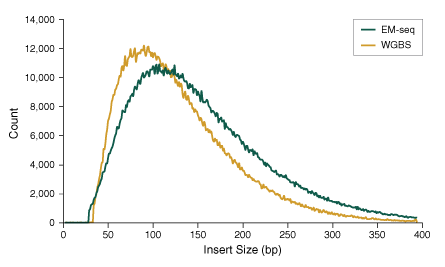 50 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold kit for bisulfite conversion. Libraries were sequenced on an Illumina MiSeq® (2 x 76 bases) and insert sizes were determined using Picard 2.18.14. The normalized frequency of each insert size was plotted, illustrating that library insert sizes are larger for EM-seq than for WGBS, and indicating that EM-seq does not damage DNA as bisulfite treatment does in WGBS.
50 ng Human NA12878 genomic DNA was sheared to 300 bp using the Covaris S2 instrument and used as input into EM-seq and WGBS protocols. For WGBS, NEBNext Ultra II DNA was used for library construction, followed by the Zymo Research EZ DNA Methylation-Gold kit for bisulfite conversion. Libraries were sequenced on an Illumina MiSeq® (2 x 76 bases) and insert sizes were determined using Picard 2.18.14. The normalized frequency of each insert size was plotted, illustrating that library insert sizes are larger for EM-seq than for WGBS, and indicating that EM-seq does not damage DNA as bisulfite treatment does in WGBS.
View additional performance data in our
technical note .
产品类别: Methylome Analysis, Epigenetic Analysis, Epigenetics Products
应用: DNA Methylation Analysis, Bisulfite Conversion, Methylated DNA Analysis Products , |
More +






